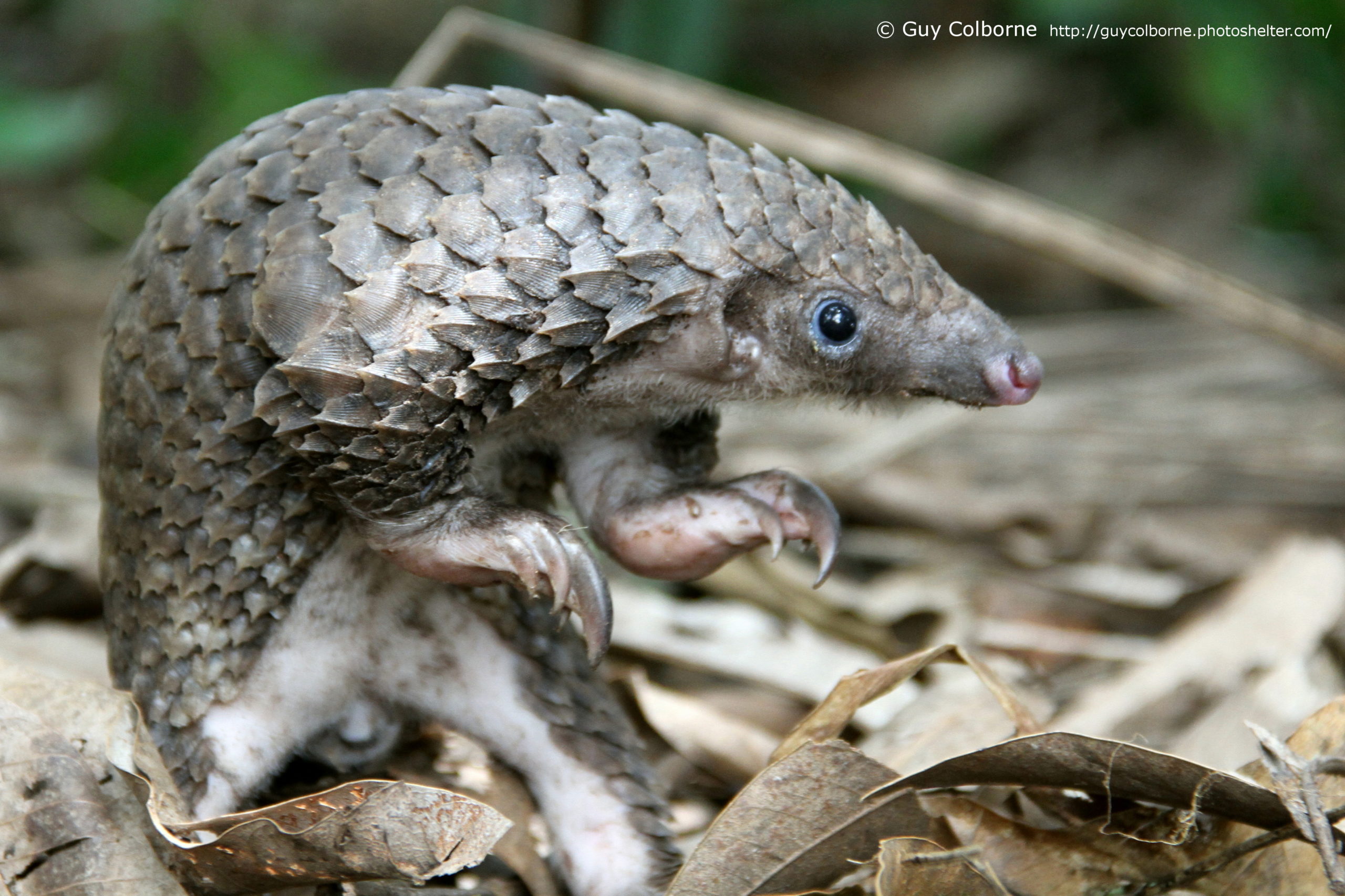
Philippe Gaubert, staff researcher at Institut de Recherche pour le Développement (IRD), Toulouse, France, gives an insight into how a better understanding of the evolutionary history of pangolins can help monitor the global pangolin trade.
The world’s most trafficked group of wild mammals is also one of the most evolutionarily unique mammalian lineages. Despite the recent international trade ban following the listing of pangolins on Appendix I of CITES, pangolins continue suffering from international trafficking, notably through elusive criminal networks that deliver a variety of pangolin forms difficult to identify by eye (e.g. isolated scales, scale powder, embryo soups).
Moreover, the lack of knowledge on pangolin evolutionary history complicates attempts to assess the impact of pangolin trafficking and to develop tools to support comprehensive management plans. With collaborators from nine countries (including Belgium, Cameroon, China, France, Ghana, Guinea, Portugal, Sri Lanka, and United States of America), a worldwide initiative was launched to establish the first, comprehensive phylogenetic framework (species tree) of pangolins.
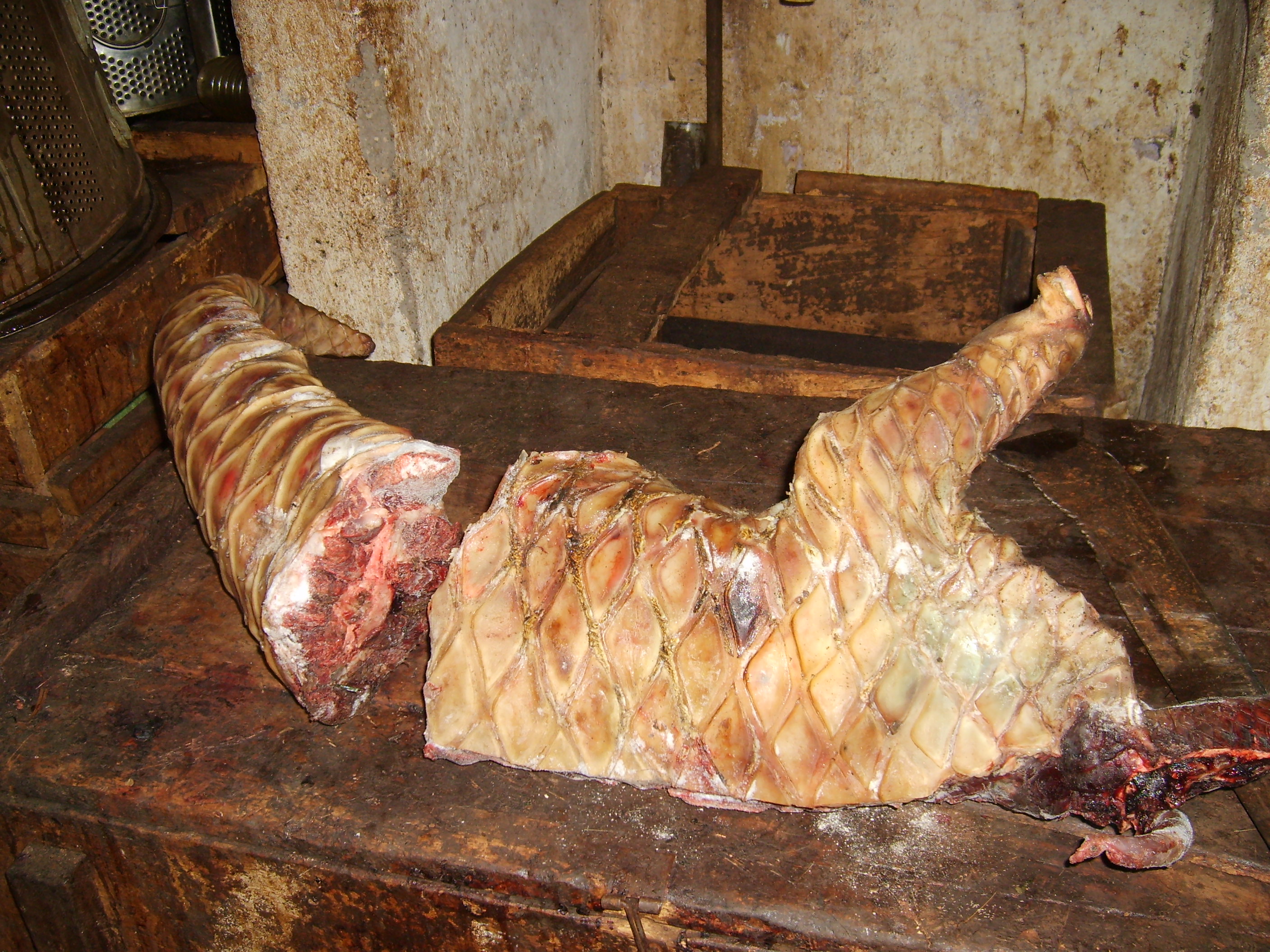

Processed carcass of a giant pangolin sold at a bushmeat market in Cameroon. Photo Credit: Flobert Njiokou.
The project involved obtaining genetic samples from pangolins sourced primarily from bushmeat markets. Where it was difficult to obtain samples, the list was completed by assessing museum specimens and their degraded DNA, as was the case with the small population of the Philippine pangolin. The ultimate objective of our initiative was to revise pangolin taxonomic boundaries and to identify the most suitable genetic markers for the molecular tracking of worldwide pangolin trafficking.
Combining mitogenomic and nuclear gene datasets, we first estimated that extant pangolins were an ancient mammalian lineage of Upper Eocene – Oligocene origin (ca. 31–45 million years ago), splitting into Asian and African lineages before the Oligocene-Miocene boundary (23 million years ago).
Our taxonomic analyses based on genetic and morphological data (i) supported the designation of a new subfamily of small African pangolins, the Phatagininae and (ii) revealed an important level of cryptic –previously undetected– genetic diversity within the wide-ranging African common (or white-bellied) pangolin, suggesting that new species of pangolins remain to be described.
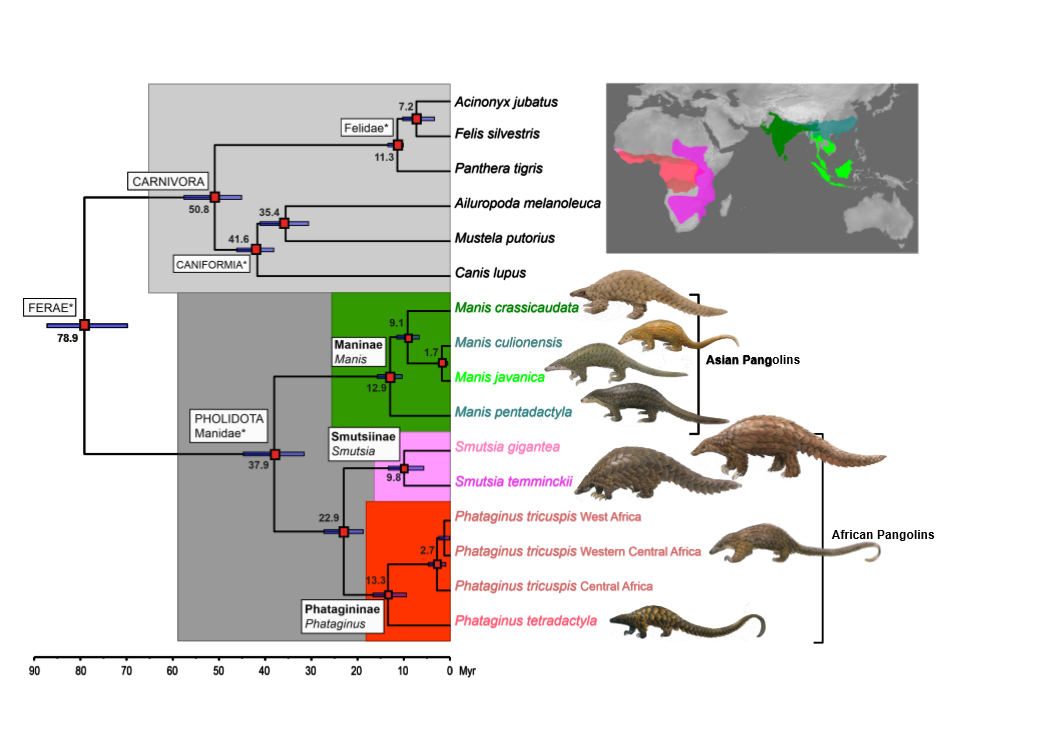
Time-calibrated, molecular phylogenetic tree of pangolins, summarizing their distribution and revised classification. Time to most recent common ancestors (in million years) are indicated at the tree nodes. Modified from Gaubert et al. (2017).
Importantly, we set up a new generation of molecular tools (markers) that should improve our ability to trace the worldwide pangolin trade. These markers included full mitogenomes, which are ‘super markers’ providing > 5,000 diagnostic sites among pangolins’ species and lineages. We also designed more traditional and affordable approaches that targeted diagnostic portions of specific genes, notably from the nuclear genome. These genetic markers allow the identification of all eight species of pangolins, and of geographic lineages within species with large distributions such as the African common (or white-bellied) pangolin, which will be useful to efficiently track pangolin trafficking worldwide. To date, only a few studies have used genetic tools to explore the geographic structure of pangolins and trace their seizures. Our results will facilitate more in-depth analyses of geographic subdivisions in the pangolins’ ranges and will hopefully help monitor pangolin trade worldwide, providing crucial information to wildlife trade groups.
A better understanding of the evolutionary history of pangolins could help to save them from extinction.
Philippe Gaubert is a staff researcher at Institut de Recherche pour le Développement (IRD), Laboratoire Evolution et Diversité Biologique, Université Paul Sabatier, Toulouse, France, studying the evolution, biogeography and conservation genetics of Ferae (Carnivores and Pangolins). He is first author to ‘“The complete phylogeny of pangolins: scaling up resources for the molecular tracing of the most trafficked mammals on Earth”, recently published in Journal of Heridity.



